| Day | Date | Time | Title |
|---|---|---|---|
| 1 | 2026-05-07 | 09:00 - 09:30 | Welcome and Introduction to Research Data Management |
| 1 | 2026-05-07 | 09:30 - 10:30 | Project & Data Organization |
| 1 | 2026-05-07 | 10:30 - 11:30 | Data Management Plans (DMPs) |
| 1 | 2026-05-07 | 11:30 - 12:30 | Lunch Break |
| 1 | 2026-05-07 | 12:30 - 13:30 | Command Line |
| 1 | 2026-05-07 | 13:30 - 14:30 | Best practices for rectangular data |
| 1 | 2026-05-07 | 14:30 - 16:00 | Brain Imaging Data Structure (BIDS) |
Brain Imaging Data Structure (BIDS)
Research Data Management for Psychology and Neuroscience
Course at Julius-Maximilians-Universität Würzburg, RTG 2660: Approach-Avoidance
Slides | Source
14:30
Schedule
This session: Brain Imaging Data Structure (BIDS)
Objectives
💡 You understand what BIDS (Brain Imaging Data Structure) is and why it’s important for neuroimaging.
💡 You can explain the core principles of BIDS and how it solves common data organization problems.
💡 You can organize neuroimaging data according to BIDS directory structure standards.
💡 You understand the role of JSON metadata files and TSV data files in BIDS datasets.
💡 You know how to validate BIDS datasets using the BIDS validator.
💡 You understand the benefits of using BIDS for collaboration, reproducibility, and data sharing.
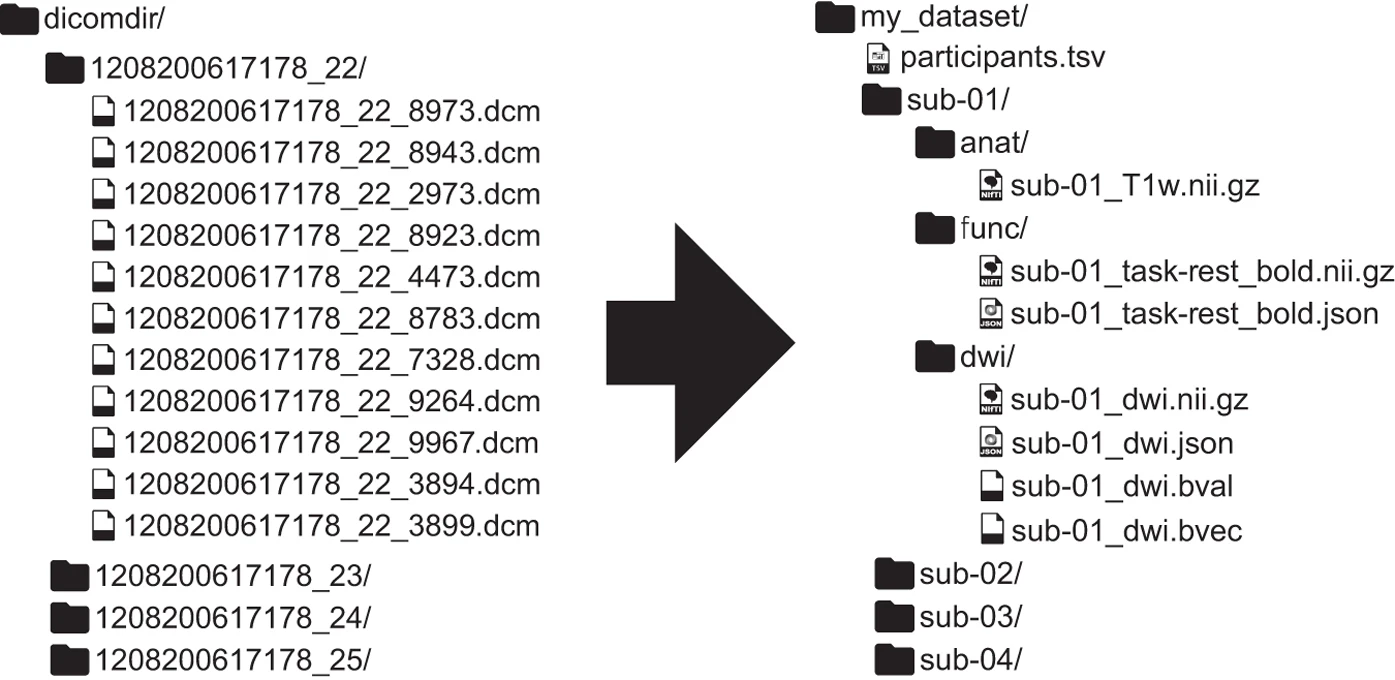
1 Introduction to BIDS
What is BIDS?
The Brain Imaging Data Structure (BIDS) is a standard to organize and describe neuroimaging and behavioral data.

Key characteristics:
- Standardized file organization and naming conventions
- Community-driven specification
- Modality-agnostic (fMRI, EEG, MEG, PET, behavior)
- Human and machine readable
- Growing ecosystem of compatible tools
The Problem BIDS Solves
Before BIDS 😞
- Every lab organized data differently
- Time wasted understanding data structure
- Difficult collaboration and data sharing
- Scripts broke when data moved
- No common metadata standards
With BIDS 😊
- Standard organization across labs
- Immediate understanding of data structure
- Easy collaboration and sharing
- Tool interoperability
- Structured metadata
Standards …
… but probably different with BIDS?

2 BIDS Principles
Core BIDS Principles
- Simplicity: Based on common file formats (NIfTI, JSON, TSV)
- Consistency: Standardized naming and folder structure
- Flexibility: Supports multiple neuroimaging modalities
- Extensibility: Community-driven extensions (BEPs)
- Validation: Tools to check compliance
- Documentation: Self-documenting through metadata files
File Naming Convention
BIDS uses entities to create standardized file names:
sub-<label>[_ses-<label>][_task-<label>][_acq-<label>][_run-<label>]_<suffix>.<ext>Examples:
sub-01_task-rest_bold.nii.gz(fMRI resting-state)sub-02_ses-001_task-faces_run-1_bold.nii.gz(task fMRI)sub-01_T1w.nii.gz(structural MRI)sub-03_task-nback_eeg.bdf(EEG data)
3 BIDS Directory Structure
Basic BIDS Dataset Structure
my-dataset/
├── dataset_description.json
├── README
├── CHANGES
├── participants.tsv
├── participants.json
└── sub-01/
├── anat/
│ ├── sub-01_T1w.nii.gz
│ └── sub-01_T1w.json
└── func/
├── sub-01_task-rest_bold.nii.gz
├── sub-01_task-rest_bold.json
└── sub-01_task-rest_events.tsvRequired Files
Every BIDS dataset must include:
dataset_description.json- Dataset metadata (name, version, authors)README- Human-readable description- At least one subject folder (
sub-<label>/) - Proper file naming following BIDS conventions
Optional but Recommended
participants.tsv- Subject demographics and characteristicsparticipants.json- Metadata for participants fileCHANGES- Version historyLICENSE- Data usage licensecode/- Analysis scriptsderivatives/- Processed data
4 Data Types and Modalities
Supported Neuroimaging Modalities
Core modalities:
- Anatomical MRI (
anat/) - Functional MRI (
func/) - Diffusion MRI (
dwi/) - Fieldmaps (
fmap/)
Electrophysiology:
- EEG (
eeg/) - MEG (
meg/) - iEEG (
ieeg/)
Other modalities: PET, microscopy, behavioral data, genetics, physiological recordings, and more!
Example: fMRI Task Data
sub-01/
└── func/
├── sub-01_task-faces_bold.nii.gz # Brain imaging data
├── sub-01_task-faces_bold.json # Imaging parameters
├── sub-01_task-faces_events.tsv # Task events/stimuli
└── sub-01_task-faces_events.json # Events metadataKey files:
.nii.gz- Brain imaging data.json- Technical metadata (TR, TE, slice timing, etc.)_events.tsv- Task timing and conditions
5 Metadata and Documentation
JSON Metadata Files
BIDS uses JSON files to store technical parameters and metadata:
TSV Files for Tabular Data
Tab-separated values for events, participants, and other tabular data:
onset duration trial_type response_time
0.0 2.5 face 1.2
3.0 2.5 object 0.9
6.0 2.5 face 1.1- Machine-readable for analysis
- Human-readable for inspection
- Version-control friendly (plain text)
6 BIDS Ecosystem
BIDS-Compatible Tools
Data organization:
- Heudiconv
- BIDScoin
- dcm2bids
Preprocessing:
- fMRIPrep
- MRIQC
- QSIPrep
Analysis:
- Nilearn
- SPM12
- FSL
- AFNI
Data sharing:
- OpenNeuro
- BIDS Validator
BIDS Validator
Automatic validation of BIDS compliance:
- Web version: bids-standard.github.io/bids-validator/
- Command-line tool:
bids-validator /path/to/dataset - Checks: File naming, required files, metadata consistency
Benefits:
- Catch errors early
- Ensure compatibility with BIDS tools
- Prepare for data sharing
7 Getting Started with BIDS
Steps to BIDSify Your Data
- Plan ahead: Consider BIDS during data collection
- Learn the specification: Read the BIDS specification
- Use conversion tools: dcm2bids, Heudiconv, etc.
- Validate: Use BIDS validator regularly
- Document: Create comprehensive README and metadata
- Share: Consider uploading to OpenNeuro
BIDS Starter Kit
The BIDS Starter Kit provides:
- Tutorials for getting started
- Templates for common files
- Best practices and examples
- Community support and resources
Benefits of Using BIDS
- Collaboration: Easier to share and understand data
- Reproducibility: Standardized organization aids replication
- Tool compatibility: Access to BIDS-aware analysis pipelines
- Data sharing: Ready for repositories like OpenNeuro
- Future-proofing: Organized data survives lab transitions
- Quality control: Built-in validation catches errors
8 Examples
Example Dataset: Task fMRI Study
bids-dataset/
├── dataset_description.json
├── README
├── participants.tsv
├── participants.json
├── task-faces_bold.json # Task-level metadata
├── sub-01/
│ ├── anat/
│ │ ├── sub-01_T1w.nii.gz
│ │ └── sub-01_T1w.json
│ └── func/
│ ├── sub-01_task-faces_run-1_bold.nii.gz
│ ├── sub-01_task-faces_run-1_bold.json
│ ├── sub-01_task-faces_run-1_events.tsv
│ ├── sub-01_task-faces_run-2_bold.nii.gz
│ ├── sub-01_task-faces_run-2_bold.json
│ └── sub-01_task-faces_run-2_events.tsv
└── sub-02/
└── ... (similar structure)Common Challenges and Solutions
Challenge: Complex naming
- Solution: Use BIDS tools for conversion
- Start with simple cases first
Challenge: Legacy data
- Solution: Gradual conversion
- Document any deviations
Challenge: Multiple sessions
- Solution: Use session folders (
ses-<label>) - Consistent session naming
Challenge: Non-standard acquisitions
- Solution: Use acquisition entity (
acq-<label>) - Document in README
9 Extensions and Future
BIDS Extension Proposals (BEPs)
BIDS evolves through community-driven extensions:
Recent additions:
- EEG/MEG data organization
- PET imaging protocols
- Microscopy data standards
- Motion data capture
- Genetics information
The Future of BIDS
- Broader adoption across neuroscience
- New modalities being standardized
- Enhanced metadata schemas
- Integration with analysis platforms
- Machine learning dataset preparation
10 Summary
Key Takeaways
- BIDS standardizes neuroimaging data organization
- Benefits everyone: researchers, collaborators, future you
- Ecosystem growth: tools, databases, and community support
- Start now: Plan BIDS organization during data collection
- Resources available: Specification, starter kit, validator, community
- Future-proof your research data management
Next Steps
For your next neuroimaging project:
- Read the BIDS specification
- Explore the BIDS Starter Kit
- Try the BIDS validator
- Join the BIDS community
- Consider your data collection protocol
Resources
- BIDS Specification: bids-specification.readthedocs.io
- BIDS Starter Kit: bids-standard.github.io/bids-starter-kit/
- BIDS Validator: bids-standard.github.io/bids-validator/
- OpenNeuro: openneuro.org
- Community Forum: neurostars.org/tag/bids
References
- Gorgolewski, K.J., Auer, T., Calhoun, V.D., et al. (2016). The brain imaging data structure, a format for organizing and describing outputs of neuroimaging experiments. Scientific Data, 3, 160044. doi:10.1038/sdata.2016.44
11 Exercises
Exercise 1
Exercise 1: Explore BIDS datasets on OpenNeuro
Objective: Familiarize yourself with BIDS-formatted datasets and understand how standardized organization facilitates data sharing and reuse.
Instructions:
Navigate to OpenNeuro:
- Visit https://openneuro.org/
- Browse the available datasets on the homepage
Select and explore a dataset:
- Choose one dataset that interests you (e.g., neuroimaging study related to your research area)
- Click on the dataset to view its details
Examine the BIDS structure:
- Click “Browse” or “Files” to view the dataset’s file structure
- Explore the following directories (if present):
- Root level files (
dataset_description.json,README,participants.tsv) - Subject folders (
sub-01,sub-02, etc.) - Session folders (if applicable:
ses-01,ses-baseline, etc.) - Modality folders (
anat,func,dwi,fmap, etc.)
- Root level files (
Explore the dataset:
- What type of neuroimaging data does it contain?
- How many subjects are included?
- What file naming patterns do you observe?
- Are there any derivative datasets associated with it?
Compare with raw data organization:
- Think about how this standardized structure compares to typical “raw” data organization
- What advantages does this standardization provide for:
- Data sharing?
- Automated processing?
- Reproducibility?
Discussion points:
- How does the BIDS structure make it easier to understand the dataset without detailed documentation?
- What challenges might researchers face when converting their data to BIDS format?
Exercise 2
Exercise 2: Hands-on BIDS practice
Objective: Learn how to create a BIDS dataset using heudiconv and use BIDS validation.
Access the hands-on tutorial: https://lennartwittkuhn.com/rdm-bids/
Questions & Discussion
Discussion time:
- Have you encountered challenges organizing neuroimaging data?
- Which BIDS modalities are relevant to your research?
- What barriers do you see to adopting BIDS?
- How might BIDS improve your workflow?
References
RDM Course
